Publications
Thioether-mediated protein ubiquitination in constructing affinity- and activity-based ubiquitinated protein probes. Gregory Davidson, Zeinab Moafian, Amanda Sensi, Zhihao Zhuang. Nature Protocols, 2025,
Cell-Based Covalent-Capture Deubiquitinase Assay for Inhibitor Discovery. Megan N. Doleschal, Jenna Miller, Sankalp Jain, Alexey V. Zakharov, Ganesha Rai, Anton Simeonov, Bolormaa Baljinnyam, Zhihao Zhuang. ACS Pharmacol. Transl. Sci. 2024, 7, 9, 2827–2839 https://doi.org/10.1021/acsptsci.4c00331
Autoinhibition of ubiquitin-specific protease 8: insights into domain interactions and mechanisms of regulation. J Biol Chem. 2024 Aug 28:107727. doi: 10.1016/j.jbc.2024.107727.
USP11 promotes prostate cancer progression by up-regulating AR and c-Myc activity. Proc Natl Acad Sci U S A. 2024 Jul 30;121(31):e2403331121. doi: 10.1073/pnas.2403331121.
Development of an efficient, effective, and economical technology for proteome analysis. Cell Rep Methods. 2024 Jun 17;4(6):100796. doi: 10.1016/j.crmeth.2024.100796.
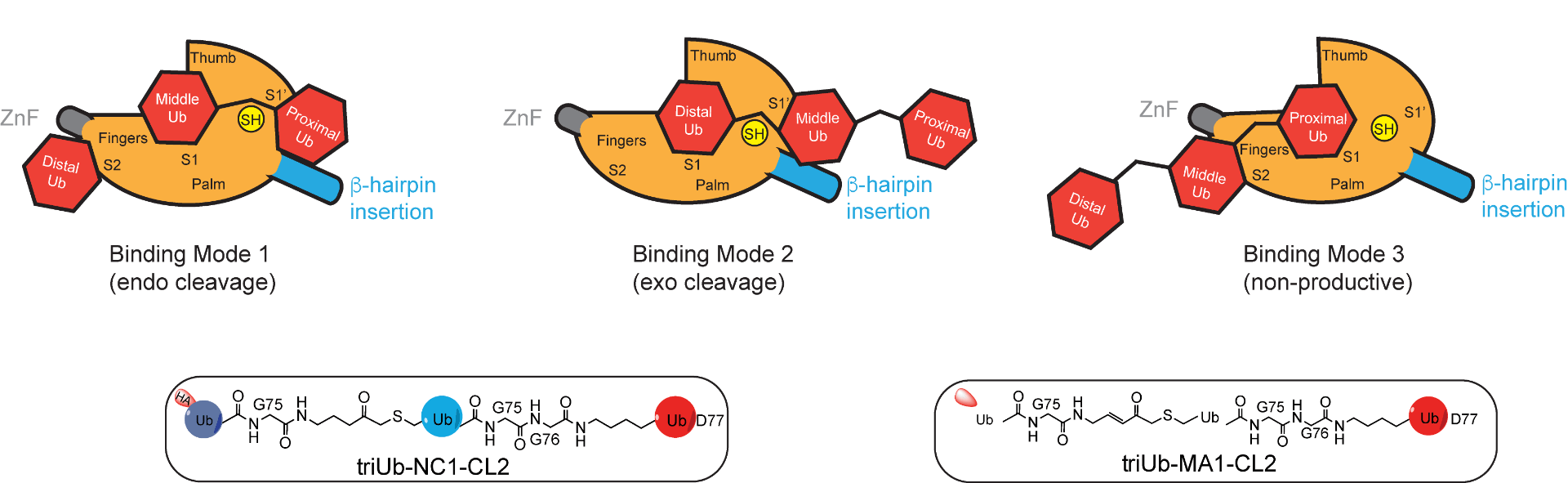
Paudel P, Banos CM, Liu Y, Zhuang Z. Triubiquitin Probes for Identification of Reader and Eraser Proteins of Branched Polyubiquitin Chains. ACS Chem Biol. 2023 Apr 21;18(4):837-847. doi: 10.1021/acschembio.2c00898. Epub 2023 Mar 27.

Shen S, Davidson GA, Yang K, Zhuang Z. Photo-activatable Ub-PCNA probes reveal new structural features of the Saccharomyces cerevisiae Polη/PCNA complex. Nucleic Acids Res. (2021) 49(16):9374-9388. doi: 10.1093/nar/gkab646. 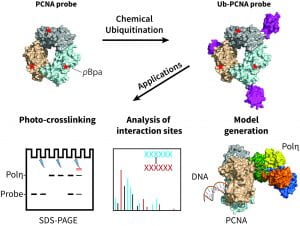
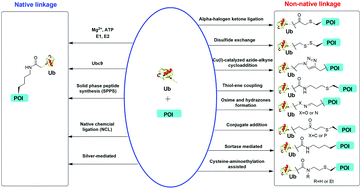
Gui W, Davidson GA, Zhuang Z. Chemical methods for protein site-specific ubiquitination RSC Chem. Biol. (2021) Advance article doi.org/10.1039/D0CB00215A
Gui W, Shen S, Zhuang Z, Photocaged Cell-Permeable Ubiquitin Probe for Temporal Profiling of Deubiquitinating Enzymes J. Am. Chem. Soc. (2020) 142(46):19493-19501 doi: 10.1021/jacs.9b12426.

Mulder MPC , Zhuang Z, Liu L , Kessler BM , Ovaa H, Editorial: Probing the Ubiquitin Landscape, Frontiers in Chemistry 2020 May 20;8:449.
Gui W, Paudel P, and Zhuang Z, Activity-Based Ubiquitin Probes for Investigation of Deubiquitinases, Comprehensive Natural Products III. 2020 : 589–602.
Paudel P, Zhang Q, Leung C, Greenberg HC, Guo Y, Chern YH, Dong A, Li Y, Vedadi M, Zhuang Z, Tong Y. Crystal structure and activity-based labeling reveal the mechanisms for linkage-specific substrate recognition by deubiquitinase USP9X PNAS (2019), 116(15):7288-7297.
Gui W, Ott CA, Yang K, Chung JS, Shen S, and Zhuang Z. Cell-Permeable Activity-Based Ubiquitin Probes Enable Intracellular Profiling of Human Deubiquitinases. J. Am. Chem. Soc. (2018), 140(39):12424.

Gong P, Davidson GA, Gui W, Yang K, Bozza WP, Zhuang Z. Activity-based ubiquitin-protein probes reveal target protein specificity of deubiquitinating enzymes. Chemical Science. (2018), 9(40):7859.
Ott C, Baljinnyam B, Zakharov A, Jadhav A, Simeonov A, Zhuang Z. Cell Lysate-Based AlphaLISA Deubiquitinase Assay Platform for Identification of Small Molecule Inhibitors. ACS Chem Biol. (2017) Sep 15;12(9):2399-2407 [PubMed]

Li G, Yuan L, Zhuang Z. Chemical Synthesis of Activity-Based Diubiquitin Probes. Methods Mol Biol. (2017);1513:223-232 [PubMed]
Tencer AH, Liang Q, Zhuang Z. Divergence in Ubiquitin Interaction and Catalysis among the Ubiquitin-Specific Protease Family Deubiquitinating Enzymes. Biochemistry (2016) Aug 23;55(33):4708-19. [PubMed]

Yang K, Li G, Gong P, Gui W, Yuan L, Zhuang Z. Chemical Protein Ubiquitylation with Preservation of the Native Cysteine Residues. ChemBioChem. (2016) Jun 2;17(11):995-8 [PubMed]

Tsutakawa SE, Yan C, Xu X, Weinacht CP, Freudenthal BD, Yang K, Zhuang Z, Washington MT, Tainer JA, Ivanov I. Structurally Distinct Ubiquitin- and Sumo-Modified PCNA: Implications for Their Distinct Roles in the DNA Damage Response. Structure. (2015) Apr 7;23(4):724-33. [PubMed]

Dexheimer TS, Rosenthal AS, Luci DK, Liang Q, Villamil MA, Chen J, Sun H, Kerns EH, Simeonov A, Jadhav A, Zhuang Z, Maloney DJ. Synthesis and structure-activity relationship studies of N-benzyl-2-phenylpyrimidin-4-amine derivatives as potent USP1/UAF1 deubiquitinase inhibitors with anticancer activity against nonsmall cell lung cancer. J Med Chem. 2014 Oct 9;57(19):8099-110 [PubMed]
Yang K, Gong P, Gokhale P, Zhuang Z. Chemical Protein Polyubiquitination Reveals the Role of a Noncanonical Polyubiquitin Chain in DNA Damage Tolerance (2014) ACS Chemical Biology, 9(8):1685-91. [PubMed]
Liang Q, Dexheimer TS, Zhang P, Rosenthal AS, Villamil MA, You C, Zhang Q, Chen J, Ott CA, Sun H, Luci DK, Yuan B, Simeonov A, Jadhav A, Xiao H, Wang Y, Maloney DJ, Zhuang Z. A selective USP1-UAF1 inhibitor links deubiquitination to DNA damage responses. (2014) Nature Chemical Biology, 10(4):298-304. Epub 2014 Feb 16 [PubMed]
Li G, Liang Q, Gong P, Tencer A and Zhuang Z. Activity-based diubiquitin probes for elucidating the linkage specificity of deubiquitinating enzymes. (2014) Chem. Comm. 50(2):216-8. Epub 2013 Nov 13. [PubMed]
Villamil M, Liang Q, and Zhuang Z. The WD40-Repeat Protein-Containing Deubiquitinase Complex: Catalysis, Regulation, and Potential for Therapeutic Intervention (2013) Cell Biochemistry and Biophysics, 67(1):111-126. [PubMed]
Richard A. Burkhart, Yu Peng, Zoë A. Norris, Renée Tholey, Vanessa A. Talbott, Qin Liang, Yongxing Ai, Kathy Miller, Shruti Lal, Joseph A. Cozzitorto, Agnieska K. Witkiewicz, Charles J. Yeo, Matthew Gehrmann, Andrew Napper, Jordan M. Winter, Janet A. Sawicki, Zhuang Z, and Jonathan R. Brody. Mitoxantrone targets human ubiquitin-specific peptidase 11 (USP11) and is a potent inhibitor of pancreatic cancer cell survival (2013) Molecular Cancer Research, 11(8):901-11. [PubMed]
Yang K, Weinacht CP, Zhuang Z. Regulatory Role of Ubiquitin in Eukaryotic DNA Translesion Synthesis. (2013), Biochemistry. 52 (19): 3217 [PubMed]
Bozza WP, Liang Q, Gong P, Zhuang Z.. Transient Kinetic Analysis of USP2-Catalyzed Deubiquitination Reveals a Conformational Rearrangement in the K48-Linked Diubiquitin Substrate (2012), Biochemistry. 51(50):10075 [PubMed]
Villamil MA, Liang Q, Chen J, Choi YS, Hou S, Lee KH, Zhuang Z. Serine Phosphorylation Is Critical for the Activation of Ubiquitin-Specific Protease 1 and Its Interaction with WD40-Repeat Protein UAF1 (2012), Biochemistry 51(45):9112 [PubMed]
Juhasz S, Balogh D, Hajdu I, Burkovics P, Villamil MA, Zhuang Z, Haracska L. Characterization of human Spartan/C1orf124, an ubiquitin-PCNA interacting regulator of DNA damage tolerance (2012), Nucleic Acids Res. 40(21):10795 [PubMed]
Song F, Thoden JB, Zhuang Z, Latham J, Trujillo M, Holden HM, Dunaway-Mariano D. The catalytic mechanism of the hotdog-fold enzyme superfamily 4-hydroxybenzoyl-CoA thioesterase from Arthrobacter sp. strain SU. (2012) Biochemistry. 51(35):7000-16 [PubMed]
Bozza WP, Yang K, Wang J, Zhuang Z. Developing peptide-based multivalent antagonists of proliferating cell nuclear antigen and a fluorescence-based PCNA binding assay (2012), Anal Biochem. 427(1):69-78 [PubMed]
Villamil MA, Chen J, Liang Q, Zhuang Z. A noncanonical cysteine protease USP1 is activated through active site modulation by USP1-associated factor 1 (2012), Biochemistry, 51(13):2829. [PubMed]
Dong J, Zhuang Z, Song F, Dunaway-Mariano D, Carey PR. A Thioester Substrate Binds to the Enzyme Arthrobacter Thioesterase in Two Ionization States; Evidence from Raman Difference Spectroscopy. (2012) J Raman Spectrosc. 43(1):65-71[PubMed]
Zhuang Z, Latham J, Song F, Zhang W, Trujillo M, Dunaway-Mariano D. Investigation of the catalytic mechanism of the hotdog-fold enzyme superfamily Pseudomonas sp. strain CBS3 4-hydroxybenzoyl-CoA thioesterase. (2012) Biochemistry. 51(3):786-94[PubMed]
Chen J, Dexheimer TS, Ai Y, Liang Q, Villamil MA, Inglese J, Maloney DJ, Jadhav A, Simeonov A, Zhuang Z. Selective and Cell-Active Inhibitors of the USP1/ UAF1 Deubiquitinase Complex Reverse Cisplatin Resistance in Non-small Cell Lung Cancer Cells. (2011) Chemistry and Biology. 18 (11): 1390. [PubMed]
Tsutakawa S, Van Wynsberghe A, Freudenthal B, Weinacht C, Gakhar L, Washington M, Zhuang Z, Tainer J, Ivanov I. Solution X-ray scattering combined with computational modeling reveals multiple conformations of covalently bound ubiquitin on PCNA. (2011) Proc. Natl. Acad. Sci. USA.108 (43):17672. [PubMed] Highlighted in Faculty 1000.
Bozza W. and Zhuang Z. Biochemical characterization of a multidomain deubiquitinating enzyme Ubp15 and the regulatory role of its terminal domains (2011) Biochemistry. 50 (29): 6423 [PubMed]
Ai Y, Wang J, Johnson RE, Haracska L, Prakash L, Zhuang Z. A novel ubiquitin binding mode in the S. cerevisiae translesion synthesis DNA polymerase η (2011) Molecular BioSystems. 7 (6): 1874. (featured as cover). [PubMed]
Chen J, Bozza W. and Zhuang Z. Ubiquitination of PCNA and its essential role in eukaryotic translesion synthesis (2011) Cell Biochemistry and Biophysics. 60: 47. [PubMed]
Chen J., Ai Y., Wang J., Haracska J., Zhuang Z. Chemically ubiquitylated PCNA as a probe for eukaryotic translesion DNA synthesis (2010) Nature Chemical Biology. 6:270. [Pubmed]
Zhuang Z, Ai Y. Processivity factor of DNA polymerase and its expanding role in normal and translesion DNA synthesis (2010) BBA-Proteins and Proteomics. 1804:1081 [PubMed]
Manosas M, Spiering MM, Zhuang Z, Benkovic SJ, Croquette V. Coupling DNA unwinding activity with primer synthesis in the bacteriophage T4 primosome. (2009) Nature Chemical Biology. 5:904. [PubMed]
Li Z, Song F, Zhuang Z, Dunaway-Mariano D, Anderson KS. Monitoring enzyme catalysis in the multimeric state: direct observation of Arthrobacter 4-hydroxybenzoyl-coenzyme A thioesterase catalytic complexes using time-resolved electrospray ionization mass spectrometry. (2009) Anal Biochem. 394:209. [PubMed]
Zhuang Z, Johnson RE, Haracska L, Prakash L, Prakash S, Benkovic SJ. Regulation of polymerase exchange between Polη and Polδ by monoubiquitination of PCNA and the movement of DNA polymerase holoenzyme. (2008) Proc. Natl. Acad. Sci. USA. 105:5361-536 [PubMed]
Zhuang Z, Song F, Zhao H, Li L, Cao J, Eisenstein E, Herzberg O and Dunaway-Mariano D. Divergence of Function in the Hotdog-Fold Enzyme Superfamily: The Bacterial Thioesterase YciA (2008) Biochemistry 47, 2789-2796
Willis MA, Zhuang Z, Song F, Howard A, Dunaway -Mariano D and Herzberg O. Structure of YciA from Haemophilus influenzae (HI0827), a hexameric broad specificity acyl-coenzyme A thioesterase. (2008) Biochemistry 47, 2797-2805
2007 and prior
Song F, Zhuang Z, Dunaway-Mariano D. Structure-activity analysis of base and enzyme-catalyzed 4-hydroxybenzoyl coenzyme A hydrolysis. (2007) Bioorg Chem. 35, 1-10.
Zhuang Z, Berdis AJ, Benkovic SJ. An alternative clamp loading pathway via the T4 clamp loader gp44/62-DNA complex. (2006) Biochemistry 45, 7976-7989.
Smiley RD, Zhuang Z, Benkovic SJ, Hammes GG. Single-molecule investigation of the T4 bacteriophage DNA polymerase holoenzyme: multiple pathways of holoenzyme formation. (2006) Biochemistry 45, 7990-7997.
Zhuang Z, Yoder BL, Burgers PM, Benkovic SJ. The structure of a ring-opened proliferating cell nuclear antigen-replication factor C complex revealed by fluorescence energy transfer. (2006) Proc. Natl. Acad. Sci. USA. 103, 2546-2551.
Song F, Zhuang Z, Finci L, Dunaway-Mariano D, Kniewel R, Buglino JA, Solorzano V, Wu J, Lima C. Structure, function and mechanism of the phenylacetate pathway hotdog-fold thioesterase PAAI. (2006) J. Biol. Chem. 281, 11028-11038.
Xi J, Zhang Z, Zhuang Z, Yang J, Spiering MM, Hammes GG, Benkovic SJ. Interaction between the T4 helicase loading protein (gp59) and the DNA polymerase (gp43): unlocking of the gp59-gp43-DNA complex to initiate assembly of a fully functional replisome. (2005) Biochemistry. 44, 7747-7756.
Yang J, Xi J, Zhuang Z, Benkovic SJ. The oligomeric T4 primase is the functional form during replication. (2005) J. Biol. Chem. 280, 25416-25423.
Xi J. Zhuang Z. Zhang Z. Selzer T. Spiering M. Hammes G. Benkovic SJ “The interaction between the T4 Helicase loading Protein (gp59) and the Polymerase (gp43): a locking mechanism to delay replication during replisome assembly.” Biochemistry (2005) 44, 2305-2318.
Willis MA, Song F, Zhuang Z, Krajewski W, Chalamasetty VR, Reddy P, Howard A, Dunaway-Mariano D, and Herzberg O. “Structure of YciI from Haemophilus influenzae (HI0828) reveals a ferredoxin-like alpha/beta-fold with a histidine/aspartate centered catalytic site.” (2005) Proteins 59, 648-652.
Zhuang Z., Spiering MM, Berdis AJ, Trakselis MA, Benkovic SJ. “ ‘Screw-cap’ clamp loader proteins that thread” (2004) Nature Structural and Molecular Biology 11, 580-581.
Yang J, Zhuang Z. Roccasecca RM, Trakselis MA, and Benkovic SJ. “The Dynamic Processivity of the T4 DNA Polymerase during Replication” (2004) Proc. Natl. Acad. Sci. USA. 101, 8289-8294.
*This paper is the subject of a commentary paper in the same issue written by Joyce CM. from Yale University, in the title of “T4 replication: what does ‘processivity’ really mean?”
Zhuang Z, Song F, Takami, H. and Dunaway-Mariano D. “The BH1999 protein of Bacillus halodurans C-125 is Gentisyl-Coenzyme A thioesterase” (2004) J. Bacteriol. 186, 393-399.
Thoden JB, Zhuang Z, Dunaway-Mariano D, Holden HM. “The structure of 4-hydroxybenzoyl-CoA thioesterase from Arthrobacter sp. strain SU.” (2003) J. Biol. Chem. 278, 43709-43716.
Zhuang Z, Gartemann KH, Eichenlaub R, Dunaway-Mariano D. “Characterization of the 4-hydroxybenzoyl-coenzyme A thioesterase from Arthrobacter sp. strain SU.” (2003) Appl Environ Microbiol. 69, 2707-2711.
Zhuang Z, Song F, Zhang W, Taylor K, Archambault A, Dunaway-Mariano D, Dong J, Carey PR. “Kinetic, Raman, NMR and Site-Directed Mutagenesis Studies of the Pseudomonas Sp. Strain CBS3 4-Hydroxybenzoyl-CoA Thioesterase Active Site” (2002) Biochemistry, 41, 11152-11160.
Thoden JB, Holden HM, Zhuang Z, Dunaway-Mariano D. “X-ray Crystallographic Analyses of Inhibitor and Substrate Complexes of Wild-type and Mutant 4-Hydroxybenzoyl CoA Thioesterase” (2002) J. Biol. Chem. 277, 27468-27476.
Zhuang Z, Song F, Martin BM, Dunaway-Mariano D. “The YbgC protein encoded by the ybgC gene of the tol-pal gene cluster of Haemophilus influenzae catalyzes acyl-coenzyme A thioester hydrolysis” (2002) FEBS Lett. 516, 161-163.